Automated grading system of retinal arterio-venous crossing patterns: A deep learning approach replicating ophthalmologist’s diagnostic process of arteriolosclerosis
Abstract
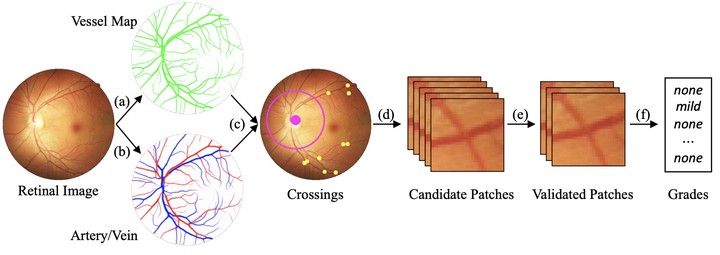
The morphological feature of retinal arterio-venous crossing patterns is a valuable source of cardiovascular risk stratification as it directly captures vascular health. Although Scheie’s classification, which was proposed in 1953, has been used to grade the severity of arteriolosclerosis as diagnostic criteria, it is not widely used in clinical settings as mastering this grading is challenging as it requires vast experience. In this paper, we propose a deep learning approach to replicate a diagnostic process of ophthalmologists while providing a checkpoint to secure explainability to understand the grading process. The proposed pipeline is three-fold to replicate a diagnostic process of ophthalmologists. First, we adopt segmentation and classification models to automatically obtain vessels in a retinal image with the corresponding artery/vein labels and find candidate arterio-venous crossing points. Second, we use a classification model to validate the true crossing point. At last, the grade of severity for the vessel crossings is classified. To better address the problem of label ambiguity and imbalanced label distribution, we propose a new model, named multi-diagnosis team network (MDTNet), in which the sub-models with different structures or different loss functions provide different decisions. MDTNet unifies these diverse theories to give the final decision with high accuracy. Our automated grading pipeline was able to validate crossing points with precision and recall of 96.3% and 96.3%, respectively. Among correctly detected crossing points, the kappa value for the agreement between the grading by a retina specialist and the estimated score was 0.85, with an accuracy of 0.92. The numerical results demonstrate that our method can achieve a good performance in both arterio-venous crossing validation and severity grading tasks following the diagnostic process of ophthalmologists. By the proposed models, we could build a pipeline reproducing ophthalmologists’ diagnostic process without requiring subjective feature extractions. The code is available (https://github.com/conscienceli/MDTNet).